Investigator, Hale Family Center For Pancreatic Cancer Research
Co-Director, Pancreatic and Biliary Tumor Center
Department Of Radiation Oncology, Dana-Farber Cancer Institute
Assistant Professor Of Radiation Oncology, Harvard Medical School
Bio Research Interests Key Publications
Bio
Dr. Joseph Mancias, MD, PhD is a radiation oncology physician-scientist at the Dana-Farber Cancer Institute and Brigham And Women’s Hospital, and assistant professor of radiation oncology at Harvard Medical School. He specializes in the care of gastrointestinal cancer patients, including pancreatic cancer patients, and runs an independent laboratory focused on the biology of pancreatic cancer.
Dr. Mancias received his medical and graduate degrees as part of the Tri-Institutional Weill Cornell/Rockefeller/Sloan-Kettering MD-PhD Program. His graduate work in the laboratory of Dr. Jonathan Goldberg at the Sloan-Kettering Institute identified molecular mechanisms of cargo export from the endoplasmic reticulum using x-ray crystallography and provided a strong foundation to study the molecular pathogenesis of disease.
His internship in internal medicine at Brigham And Women’s Hospital was followed by a residency in the Harvard Radiation Oncology Program. He subsequently completed a joint postdoctoral research fellowship in the labs of Dr. Alec Kimmelman at Dana-Farber and Dr. J. Wade Harper at Harvard Medical School, where he developed expertise in mass spectrometry-based quantitative proteomics and genetically engineered mouse models of pancreatic cancer. Dr. Mancias joined the faculty of Harvard Medical School and Dana-Farber Cancer Institute in 2016.
Research Interests
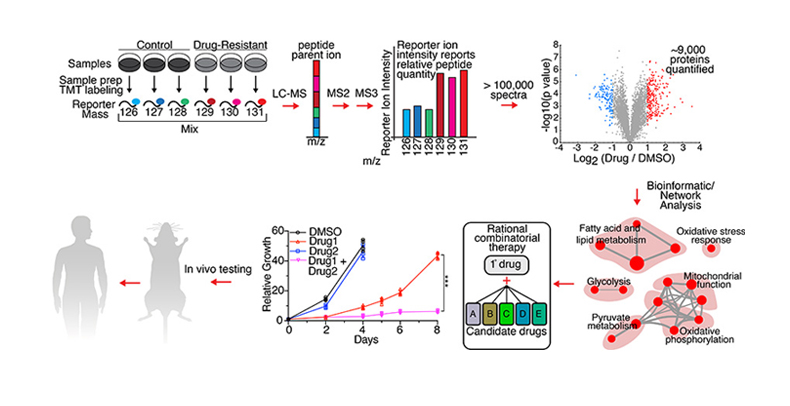
Therapeutic Resistance Quantitative Proteomics Workflow. Drug-resistant PDAC cells are profiled using Tandem Mass Tag (TMT) quantitative global proteomics. Samples are labeled with TMT reporter tags and analyzed by LC-MS (liquid chromatography-mass spectrometry), followed by MS2 and MS3 fragmentation, resulting in relative quantitation of the global proteome. A volcano plot illustrates protein abundance differences in a PDAC cell line treated with a drug. Red and Blue: top 5% up-, down-regulated proteins. Bioinformatic and network analysis leads to a map of upregulated pathways. Rational combinations are predicted and tested in vitro to determine ability to prevent or circumvent development of resistance. Effective combinations are tested in vivo in a PDAC mouse model with the eventual goal of translating to a human clinical trial for PDAC patients.
The Mancias Lab studies critical aspects of the biology of pancreatic ductal adenocarcinoma (PDAC) to develop novel therapeutic approaches.
Our research program focuses on the role of selective autophagy and therapeutic resistance in PDAC, using a comprehensive approach that combines biochemical, quantitative mass spectrometry-based proteomic, gene editing, cell biological, and mouse modeling techniques.
Recent work has shown that PDAC has a distinct dependence on autophagy (a cellular catabolic pathway) and that inhibition leads to DNA damage and metabolic defects, leading to growth suppression. The unique importance of autophagy in PDAC suggests that there may be a subset of proteins that are specifically targeted by autophagy for degradation, which would promote proliferation and survival.
To identify novel proteins degraded by autophagy in PDAC, we developed the first quantitative autophagosomal proteomics workflow. Using this methodology, we identified NCOA4 as a new selective autophagy receptor that targets the iron storage complex, ferritin, to the autophagosome for degradation, thereby releasing iron for use by the cell. Given the reliance of multiple cancers, including PDAC, on both high levels of autophagy, as well as iron, understanding the role of NCOA4 in PDAC may reveal a new therapeutic approach. To answer this question and others related to selective autophagy, we are taking an integrated biochemical, cell biologic, and in vivoapproach to answer fundamental questions of the role of NCOA4-mediated ferritin autophagy (“ferritinophagy”) in PDAC and in systemic iron homeostasis.
A second major focus of the lab is understanding the development of therapeutic resistance in PDAC using a quantitative proteomics approach. We recently showed that quantitative global proteomics of PDAC cells resistant to specific metabolic inhibitors can uncover mechanisms of resistance that can be targeted by rational combinatorial treatments. We are now adapting this quantitative proteomics platform to investigate in vitro and in vivo resistance to other targeted therapies, chemotherapy, and radiation therapy.
A final aim of the lab is to test new investigational drugs for efficacy using a genetically engineered mouse model of PDAC. This aim is part of a collaborative Hale Family Center For Pancreatic Cancer Research pre-clinical and translational effort to identify the most promising treatments for future testing in pancreatic cancer patient clinical trials.
Key Publications
Santana-Codina N, Del Rey MQ, Kapner KS, Zhang H, Gikandi A, Malcolm C, Poupault C, Kuljanin M, John KM, Biancur DE, Chen B, Das NK, Lowder KE, Hennessey CJ, Huang W, Yang A, Shah YM, Nowak JA, Aguirre AJ, Mancias JD. NCOA4-Mediated Ferritinophagy Is a Pancreatic Cancer Dependency via Maintenance of Iron Bioavailability for Iron-Sulfur Cluster Proteins. Cancer Discov. 2022 Sep 2;12(9):2180-2197. doi: 10.1158/2159-8290.CD-22-0043. PMID: 35771492; PMCID: PMC9437572.
Santana-Codina N, Chandhoke AS, Yu Q, Małachowska B, Kuljanin M, Gikandi A, Stańczak M, Gableske S, Jedrychowski MP, Scott DA, Aguirre AJ, Fendler W, Gray NS, Mancias JD. Defining and Targeting Adaptations to Oncogenic KRASG12C Inhibition Using Quantitative Temporal Proteomics. Cell Rep. 2020 Mar 31;30(13):4584-4599.e4. doi: 10.1016/j.celrep.2020.03.021. PMID: 32234489.
Santana-Codina N, Gableske S, Quiles del Rey M, Małachowska B, Jedrychowski MP, Biancur DE, Schmidt PJ, Fleming MD, Fendler W, Harper JW, Kimmelman AC, Mancias JD. NCOA4 maintains murine erythropoiesis via cell autonomous and non-autonomous mechanisms. Haematologica. 2019 Jul;104(7):1342-1354. doi: 10.3324/haematol.2018.204123. Epub 2019 Jan 10. PMID: 30630985; PMCID: PMC6601094.
Biancur DE, Paulo JA, Małachowska B, Quiles Del Rey M, Sousa CM, Wang X, Sohn ASW, Chu GC, Gygi SP, Harper JW, Fendler W, Mancias JD, Kimmelman AC. Compensatory metabolic networks in pancreatic cancers upon perturbation of glutamine metabolism. Nat Commun. 2017 Jul 3;8:15965. doi: 10.1038/ncomms15965. PMID: 28671190; PMCID: PMC5500878.
Mancias JD, Pontano Vaites L, Nissim S, Biancur DE, Kim AJ, Wang X, Liu Y, Goessling W, Kimmelman AC, Harper JW. Ferritinophagy via NCOA4 is required for erythropoiesis and is regulated by iron dependent HERC2-mediated proteolysis. Elife. 2015 Oct 5;4:e10308. doi: 10.7554/eLife.10308. PMID: 26436293; PMCID: PMC4592949.
Mancias JD, Wang X, Gygi SP, Harper JW, Kimmelman AC. Quantitative proteomics identifies NCOA4 as the cargo receptor mediating ferritinophagy. Nature. 2014 May 1;509(7498):105-9. doi: 10.1038/nature13148. Epub 2014 Mar 30. PMID: 24695223; PMCID: PMC4180099.
Santana-Codina N, Del Rey MQ, Kapner KS, Zhang H, Gikandi A, Malcolm C, Poupault C, Kuljanin M, John KM, Biancur DE, Chen B, Das NK, Lowder KE, Hennessey CJ, Huang W, Yang A, Shah YM, Nowak JA, Aguirre AJ, Mancias JD. NCOA4-Mediated Ferritinophagy Is a Pancreatic Cancer Dependency via Maintenance of Iron Bioavailability for Iron-Sulfur Cluster Proteins. Cancer Discov. 2022 Sep 2;12(9):2180-2197. doi: 10.1158/2159-8290.CD-22-0043. PMID: 35771492; PMCID: PMC9437572.
Santana-Codina N, Chandhoke AS, Yu Q, Małachowska B, Kuljanin M, Gikandi A, Stańczak M, Gableske S, Jedrychowski MP, Scott DA, Aguirre AJ, Fendler W, Gray NS, Mancias JD. Defining and Targeting Adaptations to Oncogenic KRASG12C Inhibition Using Quantitative Temporal Proteomics. Cell Rep. 2020 Mar 31;30(13):4584-4599.e4. doi: 10.1016/j.celrep.2020.03.021. PMID: 32234489.
Santana-Codina N, Gableske S, Quiles del Rey M, Małachowska B, Jedrychowski MP, Biancur DE, Schmidt PJ, Fleming MD, Fendler W, Harper JW, Kimmelman AC, Mancias JD. NCOA4 maintains murine erythropoiesis via cell autonomous and non-autonomous mechanisms. Haematologica. 2019 Jul;104(7):1342-1354. doi: 10.3324/haematol.2018.204123. Epub 2019 Jan 10. PMID: 30630985; PMCID: PMC6601094.
Biancur DE, Paulo JA, Małachowska B, Quiles Del Rey M, Sousa CM, Wang X, Sohn ASW, Chu GC, Gygi SP, Harper JW, Fendler W, Mancias JD, Kimmelman AC. Compensatory metabolic networks in pancreatic cancers upon perturbation of glutamine metabolism. Nat Commun. 2017 Jul 3;8:15965. doi: 10.1038/ncomms15965. PMID: 28671190; PMCID: PMC5500878.
Mancias JD, Pontano Vaites L, Nissim S, Biancur DE, Kim AJ, Wang X, Liu Y, Goessling W, Kimmelman AC, Harper JW. Ferritinophagy via NCOA4 is required for erythropoiesis and is regulated by iron dependent HERC2-mediated proteolysis. Elife. 2015 Oct 5;4:e10308. doi: 10.7554/eLife.10308. PMID: 26436293; PMCID: PMC4592949.
Mancias JD, Wang X, Gygi SP, Harper JW, Kimmelman AC. Quantitative proteomics identifies NCOA4 as the cargo receptor mediating ferritinophagy. Nature. 2014 May 1;509(7498):105-9. doi: 10.1038/nature13148. Epub 2014 Mar 30. PMID: 24695223; PMCID: PMC4180099.
Santana-Codina N, Singh Chandhoke A, Yu Q, Małachowska B, Kuljanin M, Gikandi A, Stańczak M, Gableske S, Jedrychowski MP, Scott DA, Aguirre AJ, Fendler W, Gray NS, Mancias JD. Defining and targeting adaptations to oncogenic KRASG12C inhibition using quantitative temporal proteomics. Cell Reports 2020; 30: 4584-4599.
Perez K, Clancy TE, Mancias JD, Rosenthal MH, Wolpin BM. When, What, and Why of Perioperative Treatment of Potentially Curable Pancreatic Adenocarcinoma. J Clin Oncol. 2017;35(5):485-9. doi: 10.1200/JCO.2016.70.2134. PubMed PMID: 28029328; PMCID: PMC5455317.
Paulo JA, Mancias JD, Gygi SP. Proteome-Wide Protein Expression Profiling Across Five Pancreatic Cell Lines. Pancreas. 2017;46(5):690-8. doi: 10.1097/MPA.0000000000000800. PubMed PMID: 28375945; PMCID: PMC5398936.
de Geus SWL, Eskander MF, Kasumova GG, Ng SC, Kent TS, Mancias JD, Callery MP, Mahadevan A, Tseng JF. Stereotactic body radiotherapy for unresected pancreatic cancer: A nationwide review. Cancer. 2017;123(21):4158-67. doi: 10.1002/cncr.30856. PubMed PMID: 28708929.
Biancur DE, Paulo JA, Malachowska B, Quiles Del Rey M, Sousa CM, Wang X, Sohn ASW, Chu GC, Gygi SP, Harper JW, Fendler W, Mancias JD, Kimmelman AC. Compensatory metabolic networks in pancreatic cancers upon perturbation of glutamine metabolism. Nat Commun. 2017;8:15965. doi: 10.1038/ncomms15965. PubMed PMID: 28671190; PMCID: PMC5500878.
Mancias JD, Kimmelman AC. Mechanisms of Selective Autophagy in Normal Physiology and Cancer. J Mol Biol. 2016;428(9 Pt A):1659-80. doi: 10.1016/j.jmb.2016.02.027. PubMed PMID: 26953261; PMCID: PMC4846542.
Dagoglu N, Callery M, Moser J, Tseng J, Kent T, Bullock A, Miksad R, Mancias JD, Mahadevan A. Stereotactic Body Radiotherapy (SBRT) Reirradiation for Recurrent Pancreas Cancer. J Cancer. 2016;7(3):283-8. doi: 10.7150/jca.13295. PubMed PMID: 26918041; PMCID: PMC4747882.
Mancias JD, Pontano Vaites L, Nissim S, Biancur DE, Kim AJ, Wang X, Liu Y, Goessling W, Kimmelman AC, Harper JW. Ferritinophagy via NCOA4 is required for erythropoiesis and is regulated by iron dependent HERC2-mediated proteolysis. Elife. 2015;4. doi: 10.7554/eLife.10308. PubMed PMID: 26436293; PMCID: PMC4592949.
Mancias JD, Wang X, Gygi SP, Harper JW, Kimmelman AC. Quantitative proteomics identifies NCOA4 as the cargo receptor mediating ferritinophagy. Nature. 2014;509(7498):105-9. doi: 10.1038/nature13148. PubMed PMID: 24695223; PMCID: PMC4180099.
Bryant KL, Mancias JD, Kimmelman AC, Der CJ. KRAS: feeding pancreatic cancer proliferation. Trends Biochem Sci. 2014;39(2):91-100. doi: 10.1016/j.tibs.2013.12.004. PubMed PMID: 24388967; PMCID: PMC3955735.
Mancias JD, Kimmelman AC. Targeting autophagy addiction in cancer. Oncotarget. 2011;2(12):1302-6. doi: 10.18632/oncotarget.384. PubMed PMID: 22185891; PMCID: PMC3282086.

