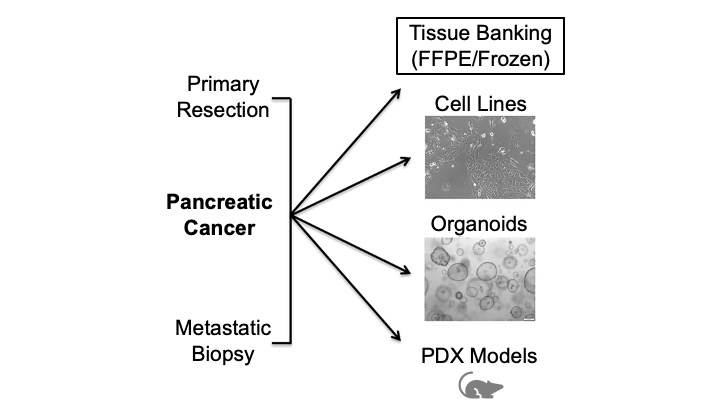
 A model system which closely mimics the in vivo pancreatic cancer tumor microenvironment in patients is necessary to make clinically-relevant discoveries and to evaluate novel therapies. The ideal model system should: represent a broad range of cancer subtypes; have an intact tumor microenvironment, including stroma, extracellular matrix, and immune cells; and be expandable at reasonable cost while still maintaining the heterogeneity and complexity of the original tumor. As no single model faithfully recapitulates the human tumor microenvironment, the Hale Family Center For Pancreatic Cancer Research employs multiple model systems to explore all facets of the pancreatic cancer niche in our studies.
A model system which closely mimics the in vivo pancreatic cancer tumor microenvironment in patients is necessary to make clinically-relevant discoveries and to evaluate novel therapies. The ideal model system should: represent a broad range of cancer subtypes; have an intact tumor microenvironment, including stroma, extracellular matrix, and immune cells; and be expandable at reasonable cost while still maintaining the heterogeneity and complexity of the original tumor. As no single model faithfully recapitulates the human tumor microenvironment, the Hale Family Center For Pancreatic Cancer Research employs multiple model systems to explore all facets of the pancreatic cancer niche in our studies.
Cellular Models
Cell Lines. Historically, the culture of pancreatic cancer cell lines involves growth of cells on plastic dishes in nutrient media. Cell lines can be grown indefinitely in culture, however, there are a limited number of available lines derived from a given cancer type and the selection of lines does not accurately reflect the genomic heterogeneity seen across the full spectrum of human cancers of that type. In addition, cell cultures lack structural organization and an intact tumor microenvironment. While specific cell lines have been favored in pancreatic cancer research (e.g., MIA PaCa2, PANC-1), there are positives and negatives associated with the use of any given cell line. The choice of cell line, therefore, depends on the questions being asked by the researcher.
Organoids. Tumor organoids consist of tumor cells that are grown in 3D on matrigel, a gelatinous protein mixture that mimics the extracellular matrix found in tissues. Typically, cells for organoids are derived from fine-needle aspiration (FNA) or surgical resection specimens. 3D culture has advantages over traditional 2D culture. The success rate of establishing organoids from patient specimens is higher than for cell lines which allows for generation of a greater diversity of lines and overall better representation of human cancer heterogeneity in an organoid biobank. The 3D system also allows for more faithful recapitulation of the spatial relationships between cells in a tumor, e.g., cell-cell and cell-matrix interactions.
Mouse Models
Patient-Derived Xenografts (PDXs). While 3D organoid culture offers some improvement over traditional 2D cell culture, organoid cultures still lack key features of the tumor microenvironment including stroma and an intact immune microenvironment. Patient-derived xenograft models (PDXs) are derived by direct implantation of tumor material from patients into immunodeficient mice which leads to tumor formation. The tumor material can be injected either subcutaneously (under the skin) or orthotopically (at the typical site of disease, i.e., injecting human pancreatic cancer cells into the pancreas of a mouse).
The advantage of PDX models is that they are thought to most closely mirror the drug responses observed in patients. These models boast an intact tumor microenvironment and preservation of elements of the immune system. Disadvantages to the use of PDXs include expense, the difficulty of generating them (e.g., lengthy passaging of cells required prior to injection and the amount of tissue needed for establishment), and limited applicability for large-scale screens.
Genetically Engineered Mouse Models (GEMMs). Genetically engineered mouse models (GEMMs) of pancreatic cancer closely mimic the human disease. The workhorse model in the Hale Center is the KPC mouse (Lox-Stop-Lox (LSL)-KrasG12D; LSL-Trp53-/-; Pdx1-cre). This mouse features combined loss of the p53 tumor suppressor gene and conversion of KRAS into its oncogenic form via KRAS G12D mutation. These alterations are present focally in pancreatic tissue since Cre expression is driven from a pancreas-specific promoter (Pdx1). Using this approach, pancreatic tumors are generated with high penetrance in a microenvironment similar to human tumors. Since immunocompetent mice are used, immune-modulatory agents (e.g., immune checkpoint inhibitors) can be tested in this system. Drawbacks to the use of GEMMs include expense and the time-consuming nature of establishing and maintaining a colony.
Modeling With The Goal Of Direct Patient Impact
With the support of a Lustgarten Foundation grant, we are collecting primary tumor specimens from patients (either from a core needle biopsy or from surgical resection) for propagation in culture. We aim to establish a “living biobank” of tumor organoids to be used as preclinical models to test both known and novel chemical compounds and/or combinations in search of promising therapies that can be brought into clinical trials for pancreatic cancer patients. We hope to establish organoids from patients and perform drug screens in real time to identify therapeuticsand/or therapeutic combinations that will be relevant for treatment of that specific patient, thus allowing us to have a direct and immediate clinical impact. 3D tumor organoids represent an intermediate model somewhere between cancer cell lines and PDXs. While this new technology holds much promise, it remains to be seen if the predictive value of organoid models is superior to other models and whether they can truly be used to identify novel diagnostic strategies for patients.

